MDAnalysis 2024 Events - Recap of the User Group Meeting and Workshops
11 Sep 2024As we near the last quarter of the year, we would like to reflect on the exciting events, including the recent MDAnalysis User Group Meeting and many workshops, that allowed us to connect with the MDAnalysis community throughout 2024.
MDAnalysis UGM (User Group Meeting)
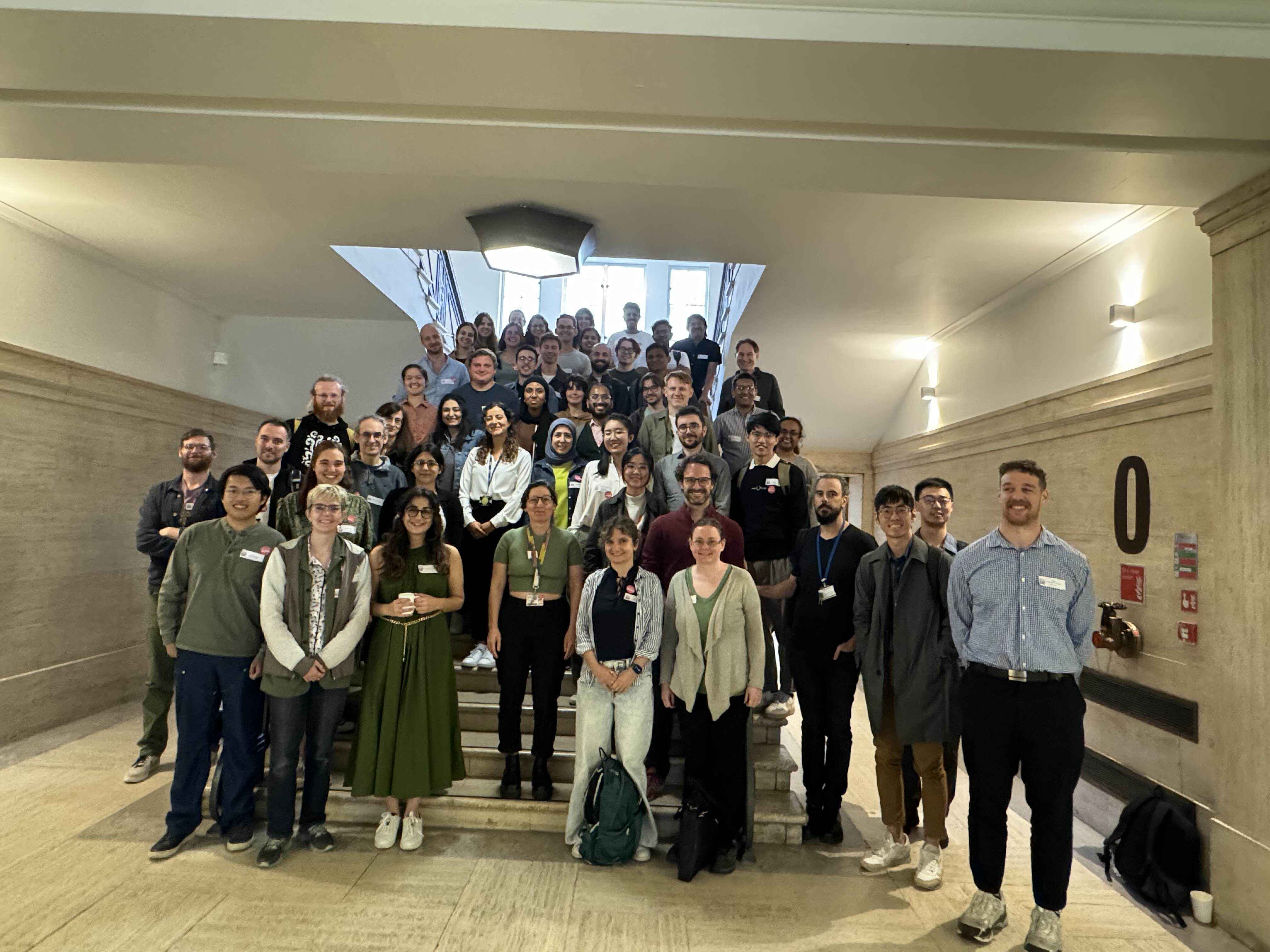

From August 21-23, 2024, 65 MDAnalysis users and developers convened in London, United Kingdom for the second MDAnalysis UGM (User Group Meeting). Hosted at King’s College London in partnership with the Thomas Young Centre, the event featured presentations and open discussions about using and developing MDAnalysis, followed by a hackathon. Talks covered a broad range of topics including, but not limited to, materials science applications, drug discovery and therapeutics, machine learning and multiscale modeling, and tools for molecular dynamics simulation analysis.
For more details, we invite you to check out the MDAnalysis/UGM2024 GitHub repository, where many of the UGM presentations and hackathon materials may be accessed.
Awards
Based on attendees’ votes, the following recognitions were awarded to UGM participants:
Best Talk
- First Place - Raquel López-Ríos de Castro
- Second Place - Lexin Chen
- Third Place - Namir Oues
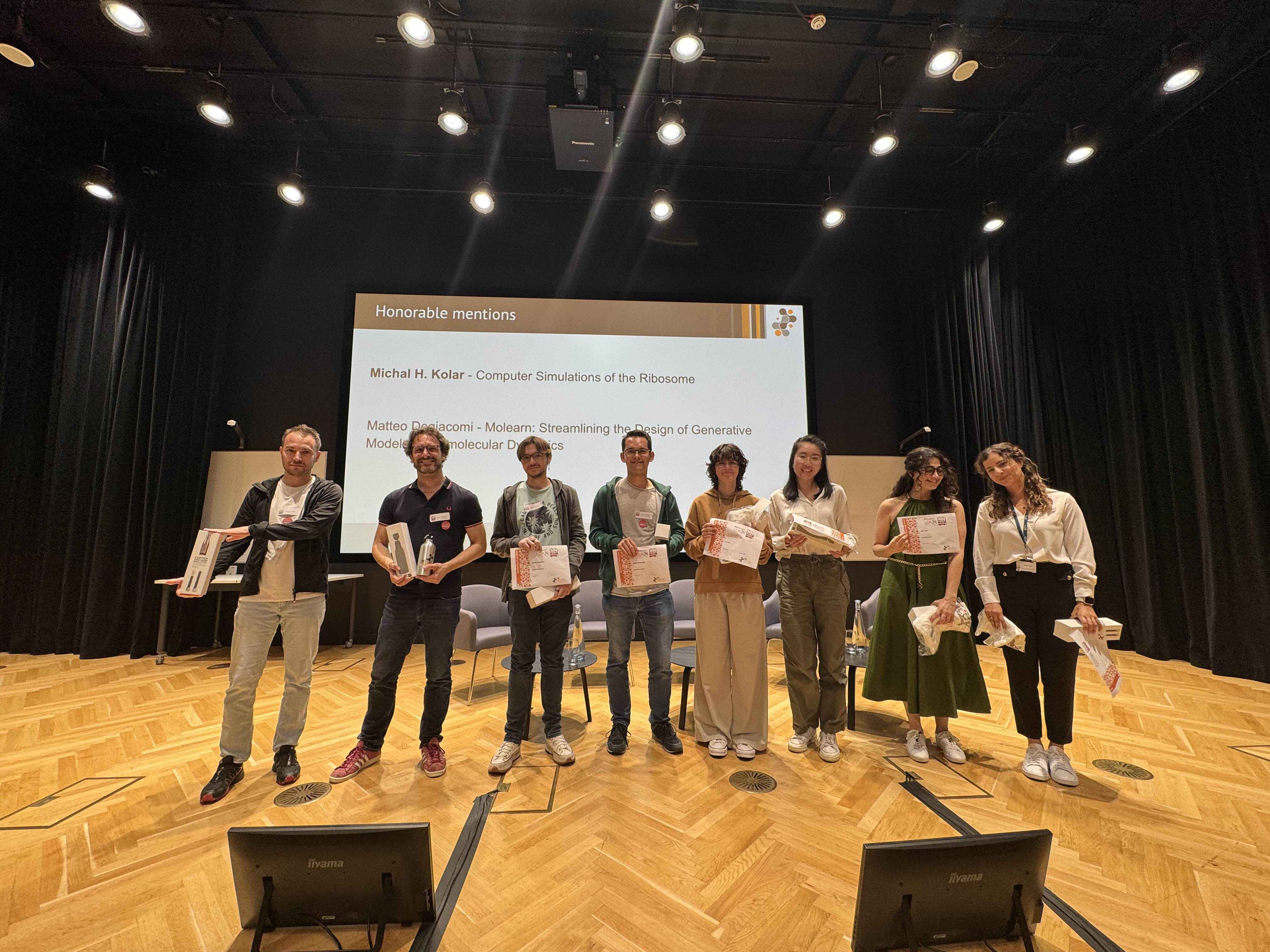
- Honorable Mentions - Michal H. Kolar, Matteo Degiacomi
Best Poster
- First Place - Asal Azar
- Second Place (TIE) - Simon Holtbrügge, Valerij Talagayev
All Star Pet (According to a pet photo contest on the #mda-pets channel on Discord)
- Pom (owned by Matteo Degiacomi)

Partners and Sponsors
We would like to thank our partners and sponsors for helping us make the 2024 MDAnalysis UGM such a success!
The Future of MDAnalysis UGMs
We would love for the MDAnalysis UGM to become an annual event. We are grateful to the Chan Zuckerberg Initiative (CZI) for supporting the 2023 and 2024 UGMs through our Essential Open Source for Science Round 5 award. However, as the 2-year grant is coming to a close, we are working hard to find ways to support potential future UGMs.
If you are interested in sponsoring, hosting at your own institution, or joining the organizing committee for any future UGMs, please get in touch with us on Discord (to join the Discord server, use the invitation link, https://discord.com/invite/fXTSfDJyxE), GitHub Discussions, or by emailing [email protected]!
MDAnalysis Training Workshops
This year was an exciting year for MDAnalysis, in that we were able to offer 4 different free online and hybrid training workshops!
Februrary 28th: Introduction to MDAnalysis and Molecular Nodes (Online)
Around 40 people joined us live on February 28, 2024 for an online workshop, during which 4 instructors (@fiona-naughton, @lilyminium, @yuxuanzhuang, @BradyAJohnston) introduced the MDAnalysis package and provided an interactive tutorial for visualizing MDAnalysis data in Molecular Nodes; the recording is available on our YouTube channel. All workshop materials and installation instructions are publicly available on the MDAnalysis/MDAnalysisWorkshop2-Intro0.5Day GitHub repository on the ‘feb24-ws’ branch.
May 10th: Introduction to Molecular Dynamics Trajectory Analysis using MDAnalysis
On May 10, 2024, 21 attendees convened in person at University College London (UCL) and another roughly 43 attendees joined online for a workshop walking participants through examples progressing from a beginner to an intermediate level. Instructors, @micaela-matta and @richardjgowers, showcased built-in analysis functions and walked participants through building custom analysis scripts; the recording of the intermediate lessons can be accessed on the MDAnalysis YouTube channel, and all workshop materials are on the MDAnalysisWorkshop-Intro1Day GitHub repo on the ‘may24-ws’ branch.

June 24th-25th: Moving from User to Developer: Analyzing Molecular Simulations and Building New Tools
In late June 2024, an MDAnalysis instructor team (@IAlibay, @orbeckst, @fiona-naughton, @yuxuanzhuang, @mikemhenry, @ianmkenney) partnered with the Molecular Sciences Software Institute (MolSSI) to offer a 2-day hybrid workshop at Arizona State University (ASU) in the Center for Biological Physics covering beginner to intermediate tutorials on the MDAnalysis library as well as tutorials on software development best practices provided by MolSSI instructors (@sjayellis, @armcdona). A total of 35 individuals attended in person, with another up to 76 attending online. All workshop materials are now publicly available in the MDAnalysisMolSSIWorkshop-Intermediate2Day GitHub repository under the ‘jun24-ws’ branch.

September 5th: MDAnalysis Intermediate Workshop, CCPBioSim Training Week
Rounding out the 2024 workshop series, @micaela-matta and @richardjgowers teamed up again for the CCPBioSim Training Week, hosted at the University of Sheffield in hybrid format, to teach participants how to build on basic MDAnalysis skills and learn about building more complex analysis scripts. There were 13 people registered for in-person attendance, and another 30 people registered for online participation. The materials for the workshop can be accessed on the MDAnalysisWorkshop-Intermediate1Day GitHub repo under the ‘ccpbiosim-24’ branch.
Acknowledgements
A special thanks goes out to the many partners and sponsors that helped us make these workshops possible:
We would like to again thank everyone who helped us make these workshops a success, including the participants, instructors, teaching assistants, and workshop organizers! If you or your organization are interested in partnering with MDAnalysis to organize future workshops, you are always welcome to fill out our Google form to discuss further.
To stay up-to-date on all MDAnalysis event offerings, be sure to follow our blog, X, LinkedIn, and Bluesky pages.